Get to know us
Our Research Areas
Fundamental molecular processes underpin the complex cellular features in large macromolecular assemblies, the intricate architecture of organelles, or the highly orchestrated contacts between organelles and cells. Cellular complexity relies on the modulation of molecule counts and molecular interactions, the control of transport across membranes, and the remodeling of membrane structures.
The next major challenge in the molecular biosciences is characterizing sub-cellular structures and their dynamics with high temporal and spatial resolution, from emergence to controlled degradation. Such understanding forms the basis for novel strategies in biomedicine, and for exciting applications such as the construction of synthetic organelles and the engineering of novel cellular functions.
Research in the IMPRS will cover the following areas of research, as well as the respective methods and experimental model system suitable to tackle our major research question, how subcellular architecture arises from molecular principles.
How cells shape membranes and regulate their content
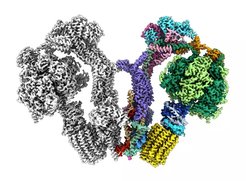
Membrane shaping is a key feature of subcellular architecture and absolutely critical for various biological processes. It is tightly regulated and occurs on largely different spatiotemporal scales. One major focus of IMPRS research topics lies on autophagy, the cellular recycling machinery for larger or membrane associated cellular waste.
It still remains a mystery how an autophagic isolation membrane suddenly arises and is shaped to specifically engulf its substrate. Đikić, Dötsch, and Münch study the molecular signals that regulate autophagy. Beck and Wilfling study the subcellular progression and autophagy pathways, while Hummer contributes the molecular modeling of those processes. Another important feature is the transport into and out of membrane-enclosed spaces that controls their size and composition. Transport of small molecules and signaling across single membranes is studied by Hänelt, Kühlbrandt and Murphy. Transport across double membranes at mitochondria, chloroplasts and the nuclear envelope are studied by Beck, Kühlbrandt, Lemke, Münch and Vonck. The architecture of these organelles is often interwoven with their molecular function. For example, ATP synthases shape membrane curvature at the tip of mitochondrial cristae, while respiratory chain complexes reside in flat adjacent membranes. Similar principles apply to chloroplast thylakoids. Building on these connections, students working synergistically in the different groups will contribute to our understanding of these processes on a larger scale.
How molecular signals shape subcellular architecture
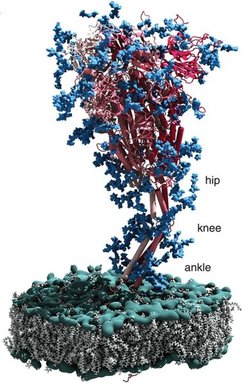
The emergence and turnover of subcellular features is spatiotemporally regulated by molecular signals. For instance, complex cellular programs have evolved to shape, monitor, and repair the Endoplasmic Reticulum (ER). Such regulatory pathways are studied by several members of the IMPRS. This includes signaling within the cytosol (Đikić, Dötsch, Knapp, Münch), but also across and within membranes (Covino, Glaubitz, Hänelt, Heilemann, Hummer, Langer), between organelles (Beck, Đikić, Wilfling) and even between cells (Acker-Palmer, Frangakis). This research topic touches upon human health relevant subjects such as the integrity of the neurovascular system (Acker-Palmer), the immune system (Schlundt), neurodegenerative diseases (Đikić, Gottschalk, Langer, Schwierz-Neumann,), host-pathogen interactions (Beck, Đikić, Hänelt, Hummer, Münch) and chromatin architecture (Kim).
How self-organizing properties contribute to biogenesis of macro-molecular assemblies
The biogenesis or turnover pathways of large macromolecular assemblies and biological membranes, such as ribosomes, nuclear pores, autophagophores and endocytic sites are another focus of the IMPRS (Beck, Frangakis, Hampölz, Wilfling).
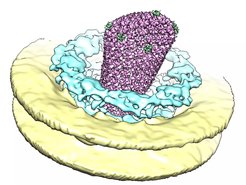
The architecture of supramolecular assemblies is studied on various different scales (Beck, Covino, Hampölz, Heilemann, Hummer, Frangakis, Kim, Kühlbrandt, Langer, Lemke, Morgner, Stelzl, Turoňová, Welsch, Wilfling) as well as the self-organizing principles that underlie the assembly of macromolecular complexes. This includes e.g. concepts from polymer physics, phase transitions and biomolecular condensates (Beck, Covino, Hampölz, Hummer, Köfinger, Lemke, Schmid, Stelzl) and is further complemented by knowledge in RNA biology and local translation that regulates the composition of membrane-less organelles and complexes (Beck, Lemke, Müller-McNicoll, Schwierz-Neumann, Schlundt, Stelzl). Taken together, this knowledge is critical to understand how the architecture of supramolecular assemblies arises from simple molecular activities encoded in their assembly pathways.
Integrative approaches based on a broad methodological repertoire
This research relies on collaborations and the integration of various different methods to allow monitoring molecular functions within life cells and to bridge across different length scales. Methods taught at the IMPRS include but are not limited to biochemistry and cell biology (Acker-Palmer, Đikić, Gottschalk, Hampölz, Müller-McNicoll), cryo-EM (Beck, Frangakis, Kühlbrandt, Murphy, Turoňová, Vonck, Welsch), NMR (Dötsch, Glaubitz, Schlundt), X-ray crystallography (Dötsch, Hänelt, Knapp), mass spectrometry (Beck, Langer, Morgner, Münch), single molecule biophysics (Heilemann, Lemke, Kim), optogenetics (Gottschalk), yeast genetics (Wilfing), animal model systems (Acker-Palmer, Gottschalk, Hampölz), life imaging and advanced light microscopy (Hampölz, Heilemann, Kim, Lemke), large scale sequencing (Müller-McNicoll), image processing (Frangakis, Turoňová) and computational modeling (Covino, Hummer, Köfinger, Schmid, Schwierz-Neumann, Stelzl).


